-Search query
-Search result
Showing all 36 items for (author: sonani & rr)

EMDB-40032: 
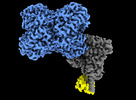
Structure of Trypanosoma (MDH)4-(Pex5)2, distal conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
EMDB-40056: 
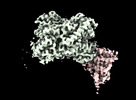
Structure of Trypanosoma docking complex
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
EMDB-40003: 
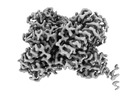
Structure of Trypanosoma (MDH)4-Pex5, close conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
EMDB-40053: 
Consensus cryo-EM reconstruction of Trypanosoma Docking complex
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-40008: 
Structure of Trypanosoma (MDH)4-PEX5, distal conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-40031: 
Structure of Trypanosoma (MDH)4-(Pex5)2, close conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-40055: 
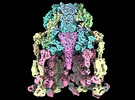
Focused map of Pex region of Trypanosoma docking complex, (MDH)4-(Pex5)1-(Pex14)1
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
EMDB-29383: 
Structure of baseplate with receptor binding complex of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH
EMDB-29353: 
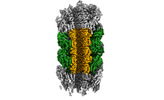
Structure of Agrobacterium tumefaciens bacteriophage Milano curved tail
Method: single particle / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-29354: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-tube
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-29355: 
Structure of Agrobacterium tumefaciens bacteriophage Milano contracted tail-sheath
Method: helical / : Sonani RR, Leiman PG, Wang F, Kreutzberger MAB, Sebastian A, Esteves NC, Kelly RJ, Scharf B, Egelman EH

EMDB-41815: 
Cryo-EM of Caulobacter crescentus Tad pilus
Method: helical / : Sonani RR, Sanchez JC, Baumgardt JK, Wright ER, Egelman EH

EMDB-41968: 
Cryo-EM of Vibrio cholera toxin co-regulated pilus (TCP)
Method: helical / : Sonani RR, Kundra S, Craig L, Egelman EH

EMDB-42279: 
Cryo-EM of Vibrio cholerae toxin co-regulated pilus - asymmetric reconstruction
Method: single particle / : Sonani RR, Kundra S, Craig L, Egelman EH
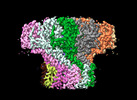
EMDB-29500: 
Portal assembly of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH
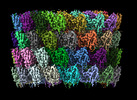
EMDB-29501: 
Collar sheath structure of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-29503: 
Neck structure of Agrobacterium phage Milano, C3 symmetry
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH
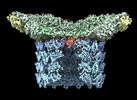
EMDB-29504: 
Structure of neck and portal vertex of Agrobacterium phage Milano, C5 symmetry
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-29512: 
Structure of tail-neck junction of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-29540: 
Structure of capsid of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH
EMDB-29541: 
Structure of neck with portal vertex of capsid of Agrobacterium phage Milano
Method: single particle / : Sonani RR, Wang F, Esteves NC, Kelly RJ, Sebastian A, Kreutzberger MAB, Leiman PG, Scharf BE, Egelman EH

EMDB-40060: 
Cryo-EM structure of Natrinema sp. J7-2 Type IV pilus
Method: helical / : Sonani RR, Kreutzberger MAB, Liu Y, Krupovic M, Egelman EH

EMDB-29215: 
Structure of the Haloferax volcanii archaeal type IV pilus
Method: helical / : Wang F, Kreutzberger MA, Baquero DP, Krupovic M, Egelman EH

EMDB-29246: 
Structure of the Saccharolobus solfataricus archaeal type IV pilus at 3 Angstrom resolution
Method: helical / : Kreutzberger MA, Wang F, Krupovic M, Egelman EH

EMDB-29247: 
Asymmetric cryo-EM structure of a curved Saccharolobus solfataricus type IV pilus
Method: single particle / : Kreutzberger MA, Wang F, Krupovic M, Egelman EH

EMDB-29249: 
Structure of the Pyrobaculum calidifontis flagellar-like archaeal type IV pilus
Method: helical / : Wang F, Kreutzberger MA, Cvirkaite-Krupovic V, Krupovic M, Egelman EH

EMDB-26995: 
Cryo-EM helical reconstruction of the EPEC H6 Curly I flagellar core
Method: helical / : Kreutzberger MAB, Chatterjee S, Frankel G, Egelman EH

EMDB-27008: 
Cryo-EM structure of the supercoiled EPEC H6 flagellar filament core Curly I waveform
Method: single particle / : Kreutzberger MA, Chatterjee S, Frankel G, Egelman EH

EMDB-27026: 
Cryo-EM structure of the supercoiled S. islandicus REY15A archaeal flagellar filament
Method: single particle / : Kreutzberger MAB, Liu J, Krupovic M, Egelman EH

EMDB-27029: 
Longer Cryo-EM Density map of the EPEC H6 Curly I bacterial flagellar filament
Method: single particle / : Kreutzberger MA, Chatterjee S, Frankel G, Egelman EH

EMDB-27059: 
Cryo-EM structure of the helical E. coli K12 flagellar filament core
Method: helical / : Sonani RR, Kreutzberger MAB, Sebastian AL, Scharf B, Egelman EH

EMDB-27060: 
Cryo-EM structure of the supercoiled E. coli K12 flagellar filament core, Normal waveform
Method: single particle / : Sonani RR, Kreutzberger MAB, Sebastian AL, Scharf B, Egelman EH

EMDB-27064: 
Cryo-EM Helical Reconstruction of the EPEC H6 flagellar filament in the Normal waveform
Method: helical / : Kreutzberger MAB, Chatterjee S, Frankel G, Egelman EH

EMDB-27065: 
Cryo-EM helical reconstruction of the S. islandicus REY15A archaeal flagellar filament
Method: helical / : Kreutzberger MAB, Liu J, Krupovic M, Egelman EH

EMDB-27076: 
Cryo-EM asymmetric reconstruction of the EPEC H6 bacterial flagellar filament Normal Waveform
Method: single particle / : Kreutzberger MAB, Chatterjee S, Frankel G, Egelman EH

EMDB-26158: 
Cryo-EM of A. pernix flagellum
Method: helical / : Wang F, Cvirkaite-Krupovic V
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
